
Welcome to
Barkoulas lab


We study development and immunity using Caenorhabditis elegans as a model organism. We are based at Imperial College London (South Kensington)
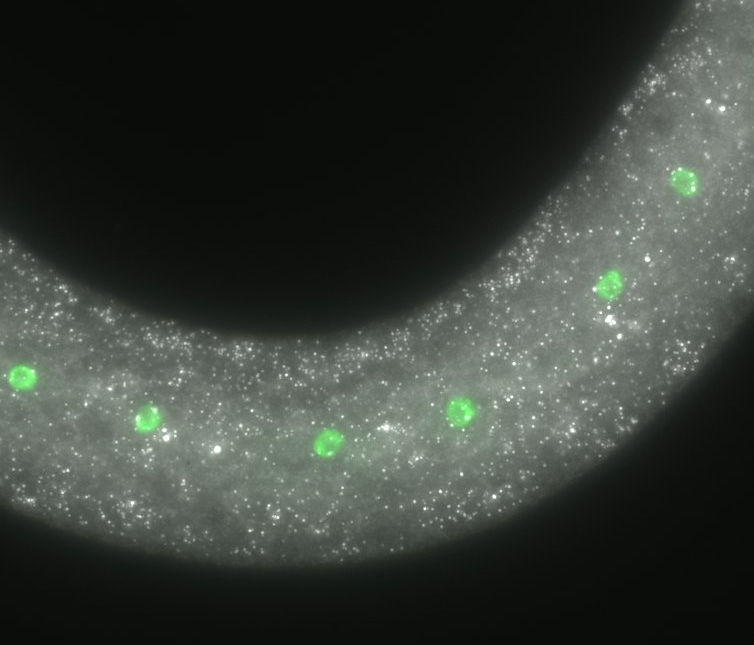
seam cells (epidermal stem cells)
Development
We study mechanisms of stem cell patterning and developmental robustness in multicellular animals using an invertebrate stem cell model.
Robustness is the ability of a system to maintain performance in the presence of perturbations and is a key principle across scientific fields. For example, engineers design systems like bridges or aeroplanes to be able to withstand a variety of perturbations and retain functionality. In a similar manner, biological systems have evolved to resist various perturbations, such as genetic mutations or changes in the environment. The model organism C. elegans is known for its stereotypical development showing an (almost) invariant pattern of cell division and differentiation events.
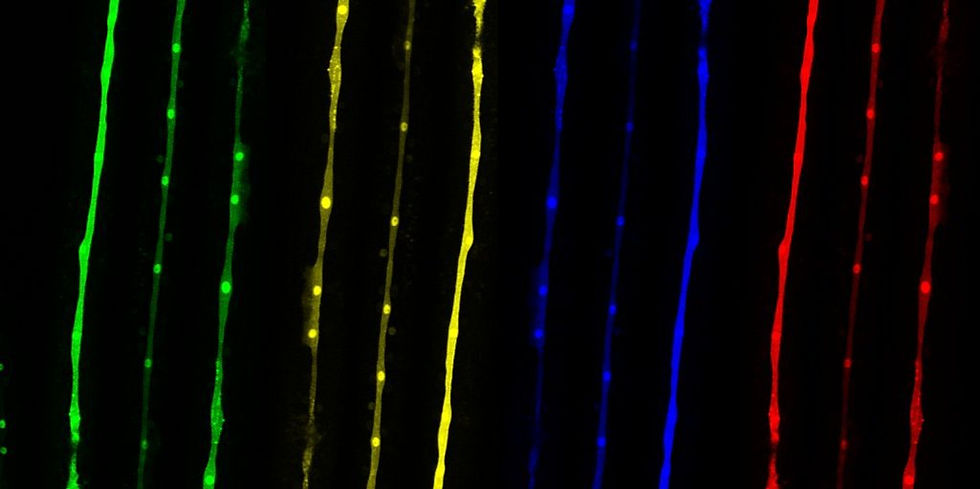
To understand the principles underlying this developmental reproducibility, we focus on a group of epidermal stem cells, known as "seam cells" (labelled here in green). These cells are found on the two lateral sides of the epidermis and show stem cell properties as they undergo symmetric cell divisions in which both daughter cells maintain the seam cell fate, or asymmetric cell divisions in which one of the daughter cells differentiates towards another cell fate. We use experimental and computational approaches to investigate the level of variation present at the molecular level among genetically identical individuals and explore the basis of the reproducibility of development in different environments and genetic backgrounds.
The key questions therefore we ask are:
i) What are the genes buffering seam cell development against stochasticity?
ii) How robust seam cell development is to environmental variation?
iii) How does genetic variation present in natural populations affect seam cell development ?
Immunity
We also study host-pathogen interactions using C. elegans as a host and oomycetes as newly discovered natural nematode pathogens.
The oomycetes represent an evolutionarily distinct lineage of eukaryotic organisms most closely related to brown algae and diatoms. The best studied oomycete species is the devastating crop pathogen Phytophthora infestans, which is infamous for causing potato blight and triggering the Great Irish famine in the 1850s. Some oomycetes infect animals as well. For example, Saprolegnia parasitica results in major fish infections in many UK rivers and is a significant threat to aquaculture. Another example is Pythium insidiosum, which infects humans among other mammals, especially in tropical and subtropical areas of the world.
So far, studies on animal infecting oomycetes have been fairly limited due to paucity of experimentally tractable model hosts. To redress this, we have established a new system to investigate animal infections by oomycetes using C. elegans as a host.
Over the last years, C. elegans has been used as a model to study mechanisms of innate immunity to various natural pathogens, such as bacteria, viruses and fungi. C. elegans does not have specialised immune cells so it relies on the first line of inducible defence, which is also known as innate immunity. We study how C. elegans senses and responds to these novel oomycete pathogens.
Contact Us
Thank you for your interest in our research. Get in touch with us if you have any questions or thoughts regarding our work.
We’d love to hear from you.
Barkoulas lab, Sir Alexander Fleming building, Imperial College London, London SW7 2AZ, UK
Where to find us:




